Ph.D. Thesis
Title Biosynthesis and Enzymology of Fluorometabolite
Production in Streptomyces Cattleya.
Adviser Prof. David O'Hagan
Thesis Committee Prof. Jim Naismith, University of St. Andrews
Scotland, UK; Prof. Tim Bugg, University of Warwick, England, UK
Essay
Organohalogens were once considered to be very rare or
even bizarre compounds, as it was believed that the few early examples
were not of natural origin but artifacts of the isolation process. With
only 30 compounds isolated in 1968 [ref. 1], the reported number in
the latest review in early 1999 [ref. 2] was 3200. Although there has
been a vast increase in the number of isolated organohalogens, only
13 of these secondary metabolites contain a fluorine atom [ref. 3].
As fluorine is the most abundant halogen in the Earth's
crust and ranks 13th in abundance of all of the elements, the small
number of isolated natural organofluorine compounds is rather surprising.
However, this can be explained by three characteristics of the fluoride
ion. Fluoride exists mainly as the tightly bound form in minerals (e.g.
fluorspar) making it highly insoluble in water and therefore restricting
its bioavailability to living organisms. Secondly, due to its high heat
of hydration, the fluoride ion is a poor nucleophile in aqueous solution
thereby limiting its participation in displacement reactions. Finally,
fluorine cannot be incorporated into organic compounds via the haloperoxidase
[ref. 4] reaction since the redox potential required for the oxidation
of fluoride is much greater than that generated by the reduction of
hydrogen peroxide.
Therefore, the mechanism by which C-F bonds are formed
in biological systems is of considerable interest. Additionally, an
understanding of the mechanism of enzymatic C-F bond formation could
provide new methods for stereospecific incorporation of fluorine into
pharmaceutically important compounds.
The aim of my thesis was to investigate the biosynthesis
of fluoroacetate and 4-fluorothreonine in the actinomycete Streptomyces
cattleya (Scheme 1) using a combination of chemical synthesis, isotopic
labelling experiments, enzymatic studies and protein purification. The
way by which fluorine is inserted into these compounds was unknown before
the research described in this thesis.
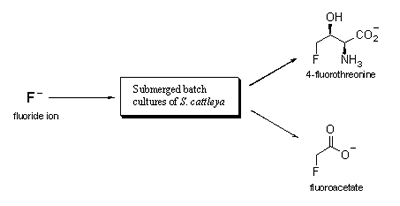
Scheme
1 Biosynthesis of 4-fluorothreonine and fluoroacetate in the bacterium
S. cattleya when batch cultures are grown in the presence of
fluoride ion [ref. 5].
Earlier work by Hamilton et al. [ref. 6] and Moss
[ref. 7] had established that fluoroacetaldehyde is the direct precursor
to fluoroacetate and that there was a biosynthetic relationship between
fluoroacetate and 4-fluorothreonine. In order to investigate the role
of fluoroacetaldehyde as the common precursor for both fluorometabolites
and to demonstrate its direct involvement in 4-fluorothreonine biosynthesis,
[1-2H]-fluoroacetaldehyde was efficiently synthesised in
three steps and subsequently administered to resting cells of S.
cattleya. Analysis of the experiment using GC-MS and 19F
NMR showed a substantial incorporation (34 %) of single 2H
into the C-3 and C-4 fragment of 4-fluorothreonine, clearly indicating
fluoroacetaldehyde as the common intermediate in the biosynthesis of
both fluorometabolites.

Scheme 2
Deuterium incorporation into 4-fluorothreonine from feeding studies
using [1-2H]-fluoroacetaldehyde.
Since the aldehyde dehydrogenase enzyme responsible for
the oxidation of fluoroacetaldehyde to fluoroacetate was already known
[ref. 8], my subsequent goal was to identify the enzyme responsible
for the transformation of fluoroacetaldehyde to 4-fluorothreonine. In
a series of experiments using cell-free extract of S. cattleya,
it was found that the second substrate for the formation of 4-fluorothreonine
is L-threonine and that the reaction is strictly
dependent on the co-factor pyridoxal phosphate (PLP). The enzyme was
partially purified and mechanistic investigations were carried out using
isotopically labelled precursors.
It was found that the discovered enzyme, threonine transaldolase,
can only accept L-threonine as substrate, thereby
representing a new class of aldolase enzyme not previously described.
All known threonine aldolases have a specificity for glycine and use
this solely with acetaldehyde to produce threonine. Furthermore, an
earlier proposal that 4-fluorothreonine is derived from glycine and
fluoroacetaldehyde5 was disproved with the discovery of the threonine
transaldolase. On the basis of these studies, a minimal mechanism for
the action of the enzyme was proposed, which is outlined in the Scheme
below.
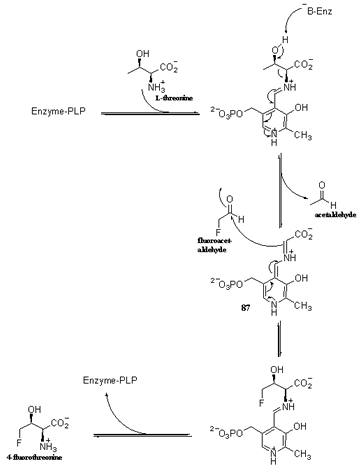
Scheme 3
Proposed pyridoxal phosphate (PLP) catalysed mechanism for the enzymatic
formation of 4-fluorothreonine from fluoroacetaldehyde and L-threonine
by threonine transaldolase.
Clearly, the main objective of my PhD was to discover
the enzyme responsible for the formation of a C-F bond. Although several
research groups have been working on this challenge for more than 40
years, there has been little progress made and proposed mechanisms for
biological fluorination are all speculative. During the course of my
PhD, I found that cell-free extract of S. cattleya, when incubated
with S-adenosyl methionine (SAM) and fluoride ion, was able to
synthesise a previously unobserved organofluorine compound, which we
proposed to be 5'-fluoro-5'-deoxyadenosine, generated by nucleophilic
attack of the fluoride ion on the C-5' carbon of SAM (Scheme 4).
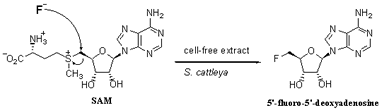
Scheme 4
Proposed mechanism for the formation of the identified organofluorine
compound.
However, isolation by semi-preparative HPLC determined
the structure to be 5'-fluoro-5'-deoxyinosine, which was found to be
a shunt product biosynthesised by the action of a deaminase on the initial
fluorination product, 5'-fluoro-5'-deoxyadeosine. The production of
5'-fluoro-5'-deoxyadenosine by the fluorinase enzyme present in S.
cattleya was confirmed by incubating cell-free extract with synthetic
5'-fluoro-5'-deoxyadenosine, which led to the accumulation of fluoroacetate
and 4-fluorothreonine.
The enzyme was efficiently purified to homogeneity from
the wildtype strain in 4 steps using ammonium sulfate precipitation,
hydrophobic interaction chromatography, gel-filtration chromatography
and anion exchange chromatography. The enzyme was found to be a hexamer
with the molecular mass of approximately 180-190 kDa.
As a potential application of the fluorinase enzyme, positron
emission tomography was investigated in collaboration with GlaxoSmithKline
and it was shown that partially purified enzyme was capable of generating
5'-[18F]-fluoro-5'-deoxyadeosine after incubation with SAM
and an aqueous solution of [18F]HF. The enzymatic radiofluorination
of an organic molecule is a novel route to the incorporation of 18F
and could have potential applications in the future.
In summary, my research gave a major insight into the
biosynthesis of fluoroacetate and 4-fluorothreonine in the bacterium
S. cattleya. Fluoroacetaldehyde was identified as the common
precursor of both fluorometabolites and the enzyme responsible for the
biotransformation of fluoroacetaldehyde to 4-fluorothreonine was identified
and isolated.
The isolation and purification of the fluorinase enzyme
represents my greatest achievement. The enzyme is the first of its class
and represents a major breakthrough in the unknown area of enzymatic
fluorination. The fluorinase enzyme now provides a system with which
to study the enzymatic syntheses of organofluorine compounds at a mechanistic
level and also opens the prospects of biotransformation routes to the
important group of fluorinated pharmaceutical compounds.
References
1. G.W. Gribble, J. Chem. Educ., 1994, 71, 907.
2. G.W. Gribble, Chem. Soc. Rev., 1999, 28, 335.
3. M. Sanada, T. Miyano, S. Iwadare, J.M. Williamson, B.H. Arison, J.L.
Smith, A.W. Douglas, J.M. Liesch and E. Inamine, J. Antibiotics,
1986, 39, 259.
4. S.L. Neidleman and J. Geigert, 'Biohalogenation: Principles, basic
roles and applications', Ellis Horwood Ltd, Chichester, 1986.
5. M. Sanada, T. Miyano, S. Iwadare, J. M. Williamson, B. H. Arison,
J. L.Smith, A. W. Douglas, J. M. Liesch and E. Inamine, J. Antibiotics,
1986, 39, 259.
6. J.T.G. Hamilton, C.D. Murphy, M.R. Amin, D. O'Hagan and D.B. Harper,
J. Chem. Soc., Perkin Trans. 1, 1998, 759.
7. S.J. Moss, PhD Thesis, 1999, University of Durham.
8. C.D. Murphy, S.J. Moss and D. O'Hagan, Appl. Environ. Microbiol.,
2001, 40, 4919.

