Australian biodiversity via its plants and marine organisms. A high-throughput
screening approach to drug discovery*
Ronald J. Quinn1,**, Priscila de Almeida Leone1, Gordon Guymer2, and
John N. A. Hooper3
1AstraZeneca R&D Griffith University, Brisbane,
Queensland, 4111, Australia; 2Queensland Herbarium, Brisbane, Queensland,
4066, Australia; 3Queensland Museum, Brisbane, Queensland, 4101, Australia
Abstract: High-throughput screening (HTS) of extracts of Australian
plants and marine organisms commenced in our laboratory in 1994. The
biota collections commenced in late 1993. The collection has in excess
of 30 000 biota samples including over 16 000 biota samples of vascular
plants, algae, and macro fungi from Queensland, and over 4000 marine
invertebrates from Australian waters. The plant collection represents
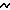 9
% of the world species diversity of higher plants, with representation
from 73 % of the world's plant families. The marine collection contains
9
% of the world species diversity of higher plants, with representation
from 73 % of the world's plant families. The marine collection contains
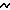 10
% of the world diversity of sponges,
10
% of the world diversity of sponges, 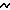 10 % of the world diversity of ascidians, and
10 % of the world diversity of ascidians, and 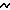 5 % of the world diversity of soft corals and gorgonians.
5 % of the world diversity of soft corals and gorgonians.
The lecture will highlight some of the advances to knowledge about
Australian biodiversity as a result of the HTS project, discuss drug
discovery using HTS, and give some examples of the chemistry arising
from the screening of the extracts.
* Lecture presented at the 3rd IUPAC International
Conference on Biodiversity (ICOB-3), Antalya, Turkey, 3-8 November 2001.
Other presentations are presented in this issue, pp. 511–584.
** Corresponding author.
Page last modified 31 May 2002.
Copyright © 2002 International Union of Pure and Applied Chemistry.
Questions or comments about IUPAC, please contact, the Secretariat.
Questions regarding the website, please contact web
manager.

